GEBRA™ Use Guide: 3bCNV – From Copy Number Variant Detection to Clinical Interpretation

In addition to SNVs, GEBRA also provides users with CNV (copy number variant) results, including deletions and duplications, detected during the analysis.
How GEBRA Detects CNVs
GEBRA uses two complementary approaches to detect CNVs.
3bCNV: Coverage Depth-Based Detection
3bCNV identifies CNVs based on depth-of-coverage signals derived from exon-level read depth in exome sequencing data.
To determine whether a genomic region represents a true copy number alteration (deletion or duplication), coverage depth is compared across a cohort of reference samples. The mean depth of coverage is calculated for each exon across the cohort, after which coverage values for individual samples are normalized to reduce inter-sample variability.
Using these normalized values, a z-score is calculated for each exon to quantify the degree of deviation from the expected coverage distribution. As a rule of thumb, 3bCNV applies a three-consecutive-exon criterion, requiring at least three adjacent exons to exhibit concordant copy-number signal shifts in order to be considered a bona fide CNV call.
This threshold significantly reduces false-positive detections arising from stochastic fluctuations in sequencing coverage.
MANTA: Breakpoint-Based Structural Variant Detection
3bCNV has certain limitations. For example, when a deletion is intragenic, the loss of an exon may be less than 50%, making it difficult for depth-based detection to capture reliably.
To overcome this limitation, MANTA is also used. MANTA is a structural variant caller that leverages breakpoint and paired-end read information. This allows the identification of smaller or partial deletions that may not be detectable solely through coverage analysis.
CNV Variant Card Structure
Let’s examine an example variant card of a heterozygous deletion identified in the 7q11.23 cytoband.
Left Panel: Basic Information
On the left-hand side of the card, the start and end positions of the called CNV are shown. The type of caller that detected this variant is also indicated accordingly.
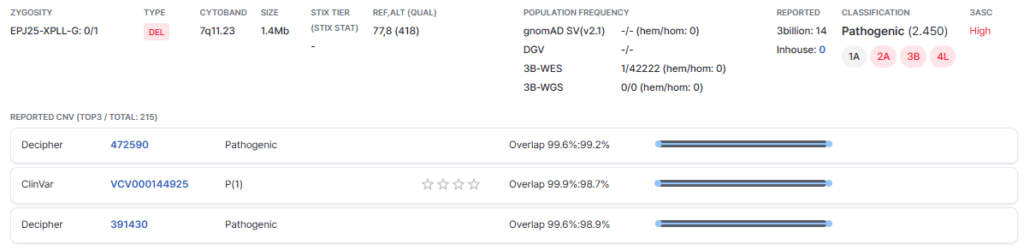
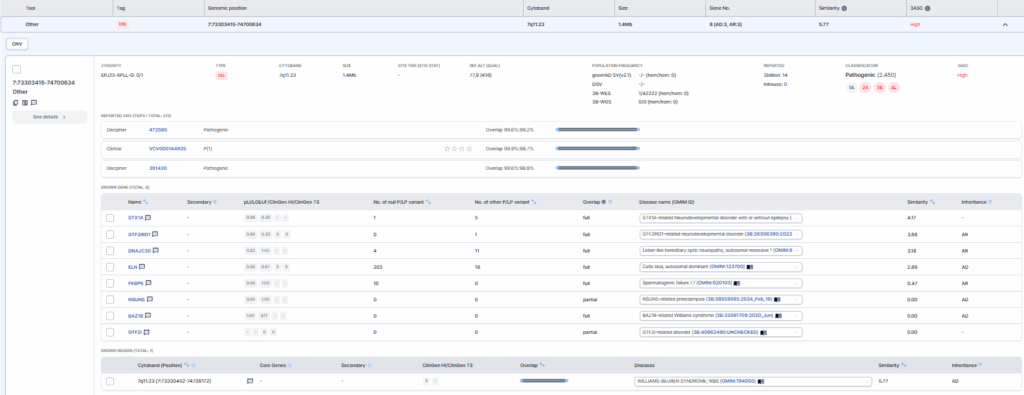
Right Panel: Detailed Annotations
Shown on the right-hand side are detailed annotations of the variant, including zygosity, cytoband, and variant size. Population frequency data indicating the occurrence of this variant in the healthy gnomAD population and the 3billion in-house database are also provided. Records of previous 3billion reports and in-house (GEBRA user) reports are displayed, along with the 3ASC class (high, mid and low) assigned to the variant.
Center Panel: Overlap Variants
In the center panel below, previously reported overlapping variants submitted to ClinVar and DECIPHER are presented. The top three variants with the highest overlap percentages with the patient’s CNV are highlighted.
Bottom Panel: Gene Information
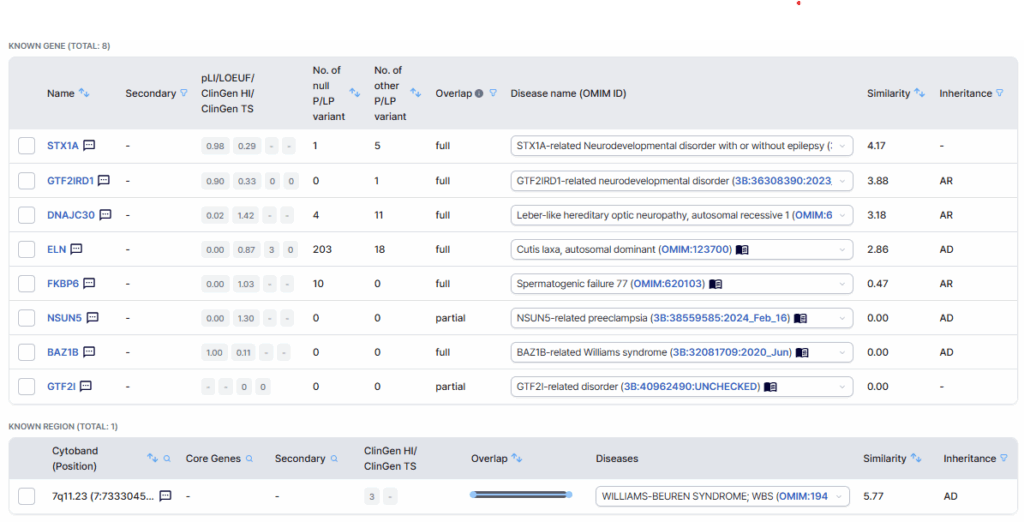
Finally, at the bottom of the card, information on genes encompassed by the detected CNV is displayed. Provided details include gene symbols, gene constraint metrics, the number and types of previously reported pathogenic/likely pathogenic (P/LP) variants per gene, associated disease entities, and phenotype similarity scores.
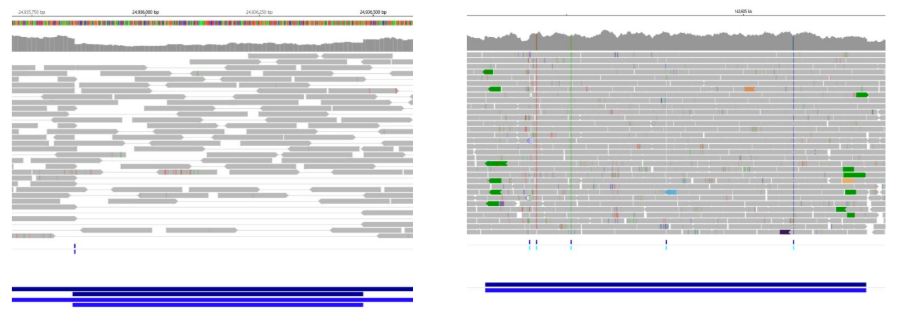
CNV Visualization
Visual representations of CNVs detected by 3bCNV are provided. The red-highlighted region at the top represents a deletion, while the green-highlighted region at the bottom represents a duplication. Each small box corresponds to an exon, with the corresponding z-score assigned for each deleted or duplicated exon.
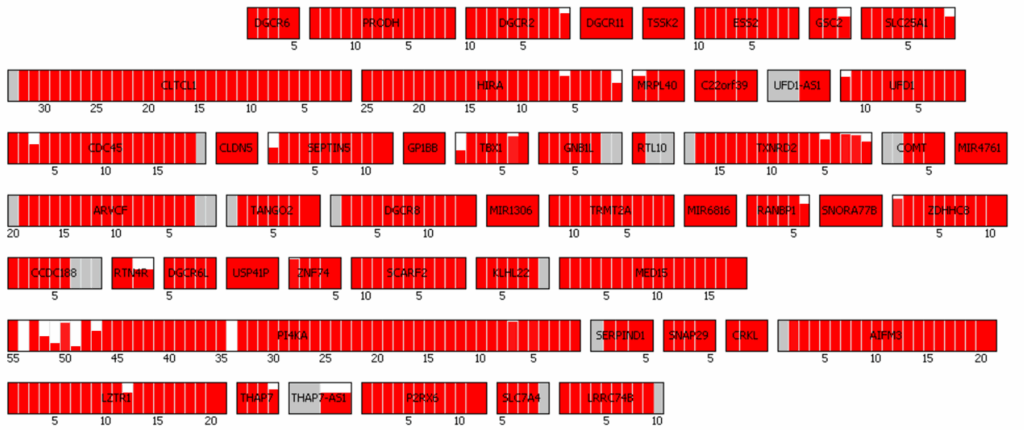
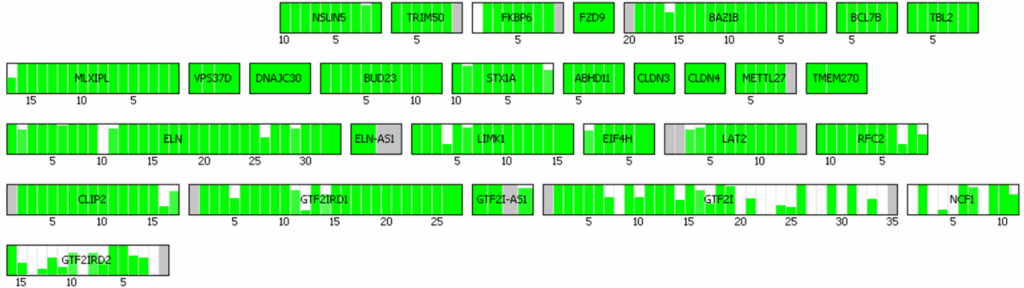
This visualization enables users to rapidly assess the overall CNV size, evaluate call confidence, and identify the genes encompassed by the CNV.
Conclusion
GEBRA provides CNV analysis results alongside SNV analysis. Using two complementary algorithms—3bCNV and MANTA—deletions and duplications are detected, and comprehensive annotation information for the detected CNVs is provided through an integrated variant card.
Frequently Asked Questions (FAQ)
Q: What is the three-consecutive-exon criterion in 3bCNV?
A: 3bCNV requires at least three adjacent exons to show concordant copy-number signal shifts to be considered a bona fide CNV call. This threshold significantly reduces false-positive detections arising from stochastic fluctuations in sequencing coverage.
Q: Why does GEBRA use both 3bCNV and MANTA?
A: 3bCNV excels at detecting large CNVs based on depth-of-coverage, but has limitations with small intragenic deletions that are smaller than 3 exons, where it’s uncertain whether they represent true deletions or duplications. MANTA complements this by leveraging breakpoint and paired-end read information to identify these small variants.
Q: How do I interpret the overlap percentages with ClinVar/DECIPHER variants?
A: The CNV variant card displays the top three variants with the highest overlap percentages. Higher overlap indicates greater relevance for referencing the pathogenicity and associated phenotypes of previously reported variants.
Q: What information does the z-score visualization provide?
A: The visualization shows each exon as a small box with its corresponding z-score. Red regions indicate deletions, while green regions indicate duplications. This allows you to rapidly assess overall CNV size, evaluate call confidence, and identify genes encompassed by the CNV.
Take a look at all GEBRA™ Related Articles:
- GEBRA™ Use Guide: 3ASC – Finding the One Causal Variant Among Thousands
- GEBRA™ Use Guide: Symptom-driven Update — When Phenotypes Evolve, Your Shortlist Should Too
- GEBRA™ Use Guide: Filters – See What Matters, Faster
- GEBRA™ Use Guide: Knowledge Base – Interpretations That Persist, Diagnoses That Compound
- GEBRA™ Use Guide: Gene Coverage – Diagnostic Confidence Starts with Coverage
- GEBRA™ Use Guide: 3bCNV – From Copy Number Variant Detection to Clinical Interpretation
- GEBRA™ Use Guide: Symptom Color Tag
Get exclusive rare disease updates
from 3billion.

3billion Inc.
3billion is dedicated to creating a world where patients with rare diseases are not neglected in diagnosis and treatment.